Compositional Perturbation Autoencoder (CPA)
Project description
CPA - Compositional Perturbation Autoencoder
What is CPA?
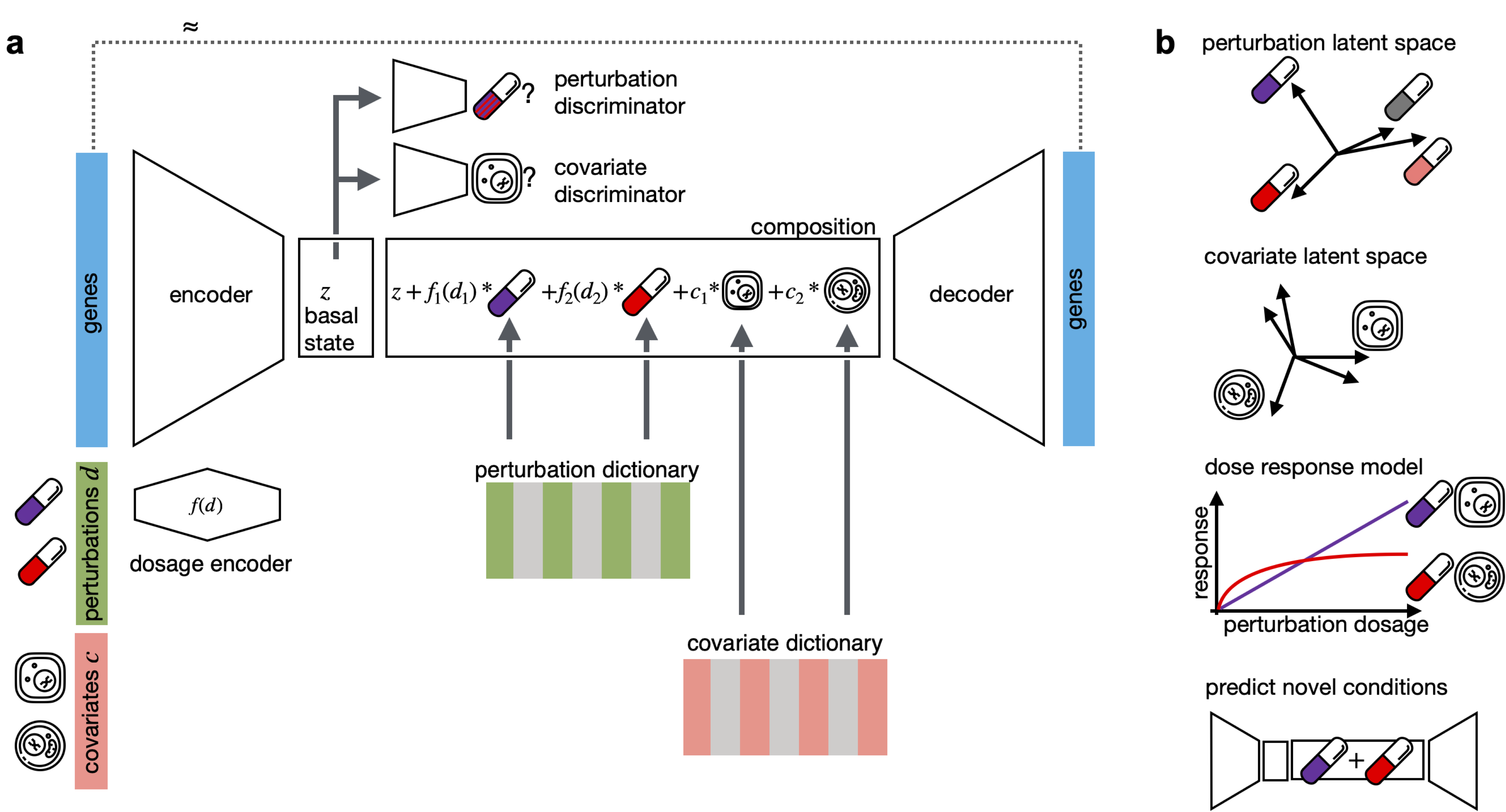
CPA is a framework to learn effects of perturbations at the single-cell level. CPA encodes and learns phenotypic drug response across different cell types, doses and drug combinations. CPA allows:
- Out-of-distribution predicitons of unseen drug combinations at various doses and among different cell types.
- Learn interpretable drug and cell type latent spaces.
- Estimate dose response curve for each perturbation and their combinations.
- Access the uncertainty of the estimations of the model.
Usage and installation
See here for documentation and tutorials.
Support and contribute
If you have a question or new architecture or a model that could be integrated into our pipeline, you can post an issue
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
cpa-tools-0.2.0.tar.gz
(35.6 kB
view details)
Built Distribution
cpa_tools-0.2.0-py3-none-any.whl
(36.5 kB
view details)
File details
Details for the file cpa-tools-0.2.0.tar.gz.
File metadata
- Download URL: cpa-tools-0.2.0.tar.gz
- Upload date:
- Size: 35.6 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: poetry/1.1.13 CPython/3.9.7 Linux/5.4.0-100-generic
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 | 6e909a23a58bd5aa4df191f737455f31d1f641513d9d08bfe981d102c7574255 |
|
| MD5 | 2d29e5fbfe19847ed7b210c37e5f0d24 |
|
| BLAKE2b-256 | 68c0bcdb19d155525f679154a639903b2e4af14010a2145b9ad133062b3c6294 |
File details
Details for the file cpa_tools-0.2.0-py3-none-any.whl.
File metadata
- Download URL: cpa_tools-0.2.0-py3-none-any.whl
- Upload date:
- Size: 36.5 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: poetry/1.1.13 CPython/3.9.7 Linux/5.4.0-100-generic
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 | 73144fc2d493c26686ea34983997a95f3fb1a4ee7c98b53a19f6cacd2af8c338 |
|
| MD5 | 53764435e1b9368d53091447927f5481 |
|
| BLAKE2b-256 | a90b3820c514343824b8fe8b6988059bef3587fe5245665671d19ce1d67b9b9f |