Compositional Perturbation Autoencoder (CPA)
Project description
CPA - Compositional Perturbation Autoencoder
What is CPA?
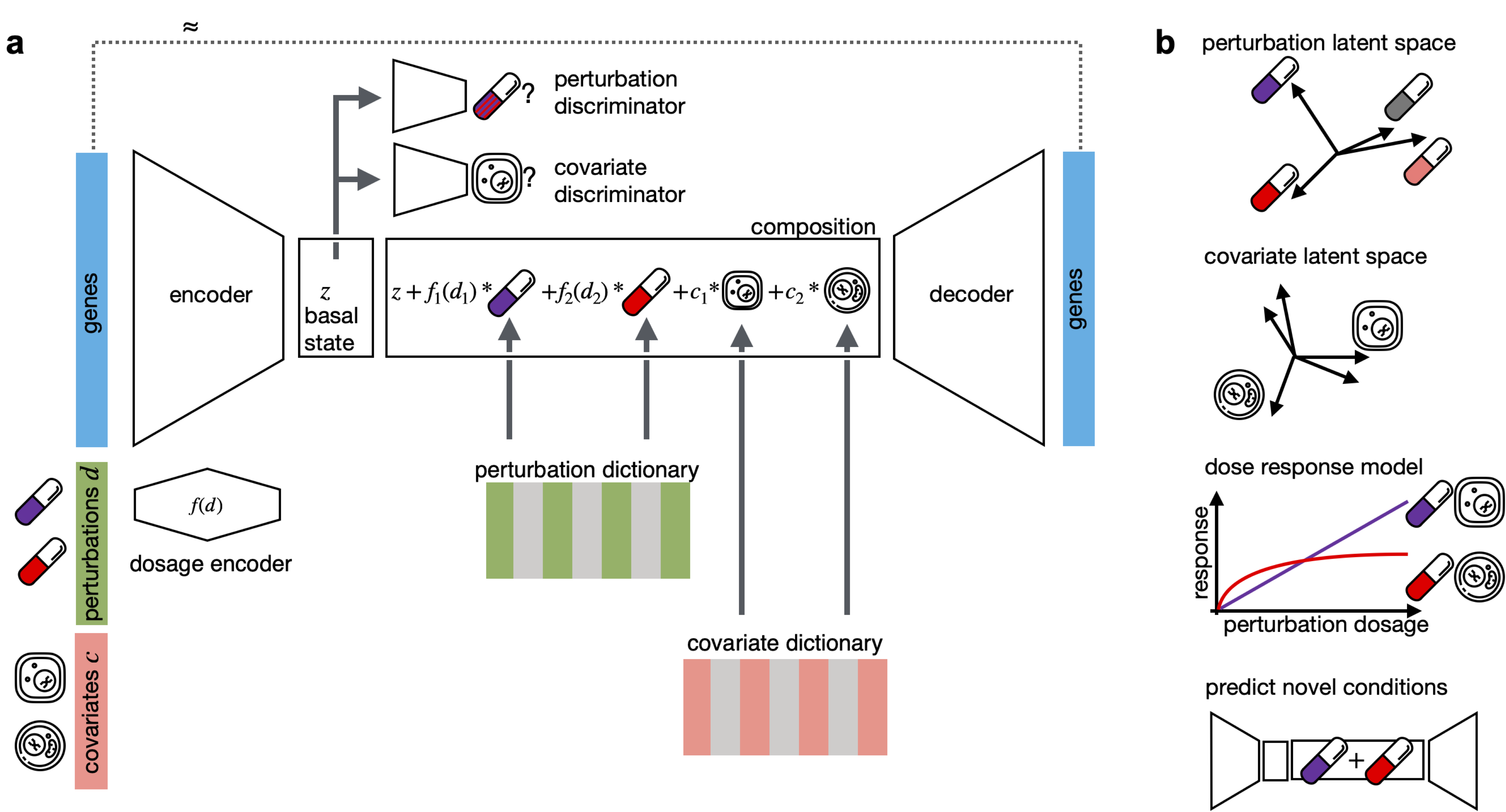
CPA is a framework to learn effects of perturbations at the single-cell level. CPA encodes and learns phenotypic drug response across different cell types, doses and drug combinations. CPA allows:
- Out-of-distribution predictions of unseen drug combinations at various doses and among different cell types.
- Learn interpretable drug and cell type latent spaces.
- Estimate dose response curve for each perturbation and their combinations.
- Access the uncertainty of the estimations of the model.
Usage and installation
See here for documentation and tutorials.
Support and contribute
If you have a question or new architecture or a model that could be integrated into our pipeline, you can post an issue
Acknowledgment
This code is inspired by an early implementatiom by Pierre Boyeau using scvi-tools.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
cpa-tools-0.3.0.tar.gz
(34.8 kB
view details)
Built Distribution
cpa_tools-0.3.0-py3-none-any.whl
(35.7 kB
view details)
File details
Details for the file cpa-tools-0.3.0.tar.gz.
File metadata
- Download URL: cpa-tools-0.3.0.tar.gz
- Upload date:
- Size: 34.8 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: poetry/1.1.13 CPython/3.8.13 Linux/5.4.0-110-generic
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 | ba46d7a07a93361e40ae9c6df93536d18339dad953e23c83d71638a131ec2244 |
|
| MD5 | 3e3c6f6f47913d20c701f12015e28815 |
|
| BLAKE2b-256 | d7f5b00621b01d4564385478c566329799cdc9e8b3f10e9e3f215c2297f9646f |
File details
Details for the file cpa_tools-0.3.0-py3-none-any.whl.
File metadata
- Download URL: cpa_tools-0.3.0-py3-none-any.whl
- Upload date:
- Size: 35.7 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: poetry/1.1.13 CPython/3.8.13 Linux/5.4.0-110-generic
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 | 11d0f98cf1448846a989c67f0756414547bb6fbace4d5161b8256f90819498a3 |
|
| MD5 | 13304fefa8ce0a8501a7502f1ed7e973 |
|
| BLAKE2b-256 | 4f9545e250aa4c512effc15ab64008b398f57feb8944b0cc7e42d408e2ec9e7e |