Compositional Perturbation Autoencoder (CPA)
Project description
CPA - Compositional Perturbation Autoencoder
What is CPA?
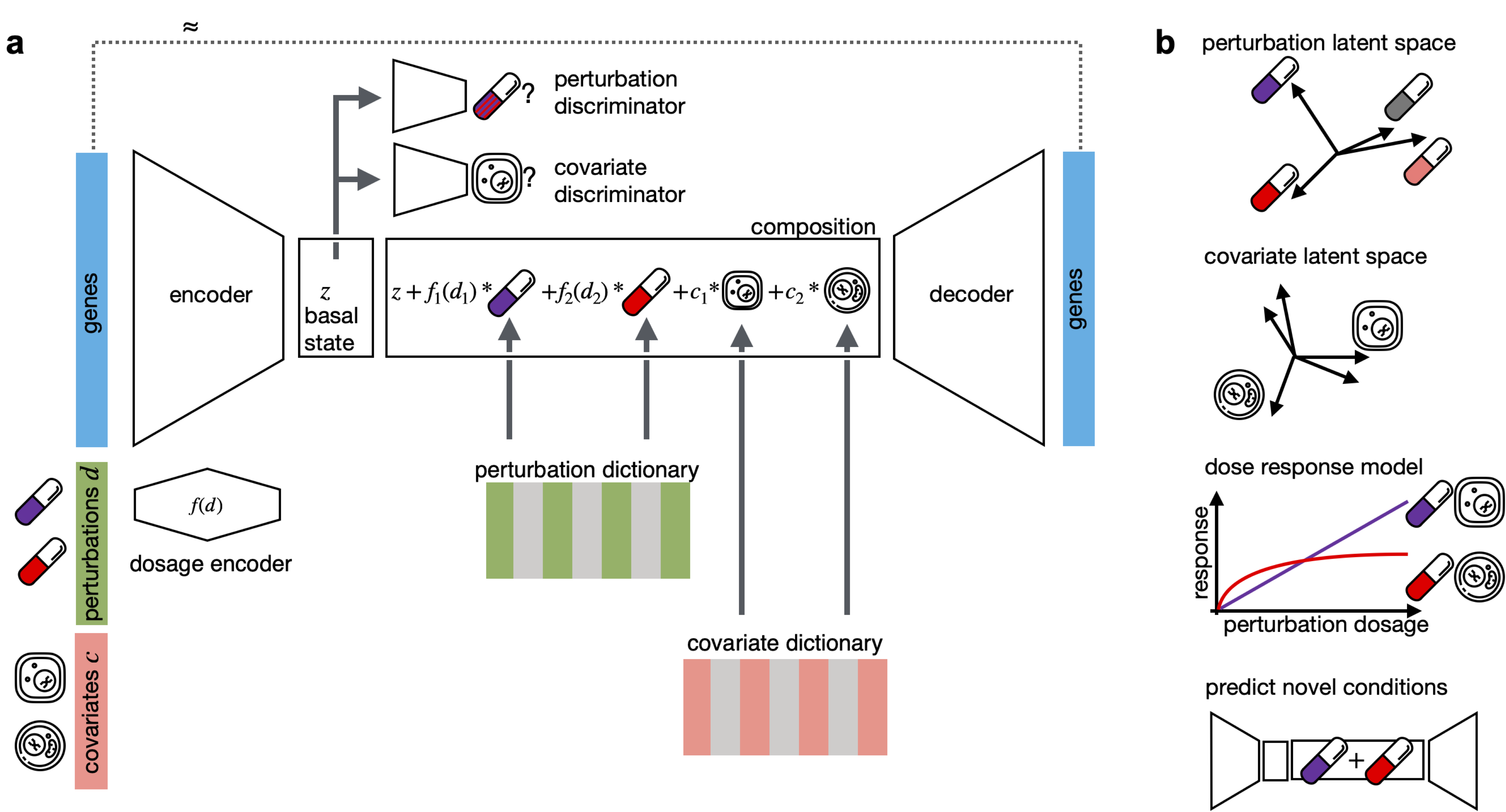
CPA is a framework to learn effects of perturbations at the single-cell level. CPA encodes and learns phenotypic drug response across different cell types, doses and drug combinations. CPA allows:
- Out-of-distribution predictions of unseen drug combinations at various doses and among different cell types.
- Learn interpretable drug and cell type latent spaces.
- Estimate dose response curve for each perturbation and their combinations.
- Access the uncertainty of the estimations of the model.
Usage and installation
See here for documentation and tutorials.
Support and contribute
If you have a question or new architecture or a model that could be integrated into our pipeline, you can post an issue
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
cpa-tools-0.4.0.tar.gz
(36.7 kB
view details)
Built Distribution
cpa_tools-0.4.0-py3-none-any.whl
(37.8 kB
view details)
File details
Details for the file cpa-tools-0.4.0.tar.gz.
File metadata
- Download URL: cpa-tools-0.4.0.tar.gz
- Upload date:
- Size: 36.7 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: python-httpx/0.25.0
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 | 802a916b5ae19e94b779128607d8deaa479851ca51e344c7dc6e33715b9733d6 |
|
| MD5 | df53d3b83c6f62a8f3a367dd468f141d |
|
| BLAKE2b-256 | c830da161ed2490be69d8cc98b76e8e20fb3216db27611fa85223e5173905372 |
File details
Details for the file cpa_tools-0.4.0-py3-none-any.whl.
File metadata
- Download URL: cpa_tools-0.4.0-py3-none-any.whl
- Upload date:
- Size: 37.8 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: python-httpx/0.25.0
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 | 8e14dabe75e8ea83547be64827a97cb374ad710202a6767b056ec8f6783efd4e |
|
| MD5 | e9794a3e044c5d8f56571dae755e98a1 |
|
| BLAKE2b-256 | b0347d6a20514e4405c56cf5708fdf01340fa1386852fc164b6f27ecb9318c7f |