Compositional Perturbation Autoencoder (CPA)
Project description
CPA - Compositional Perturbation Autoencoder
What is CPA?
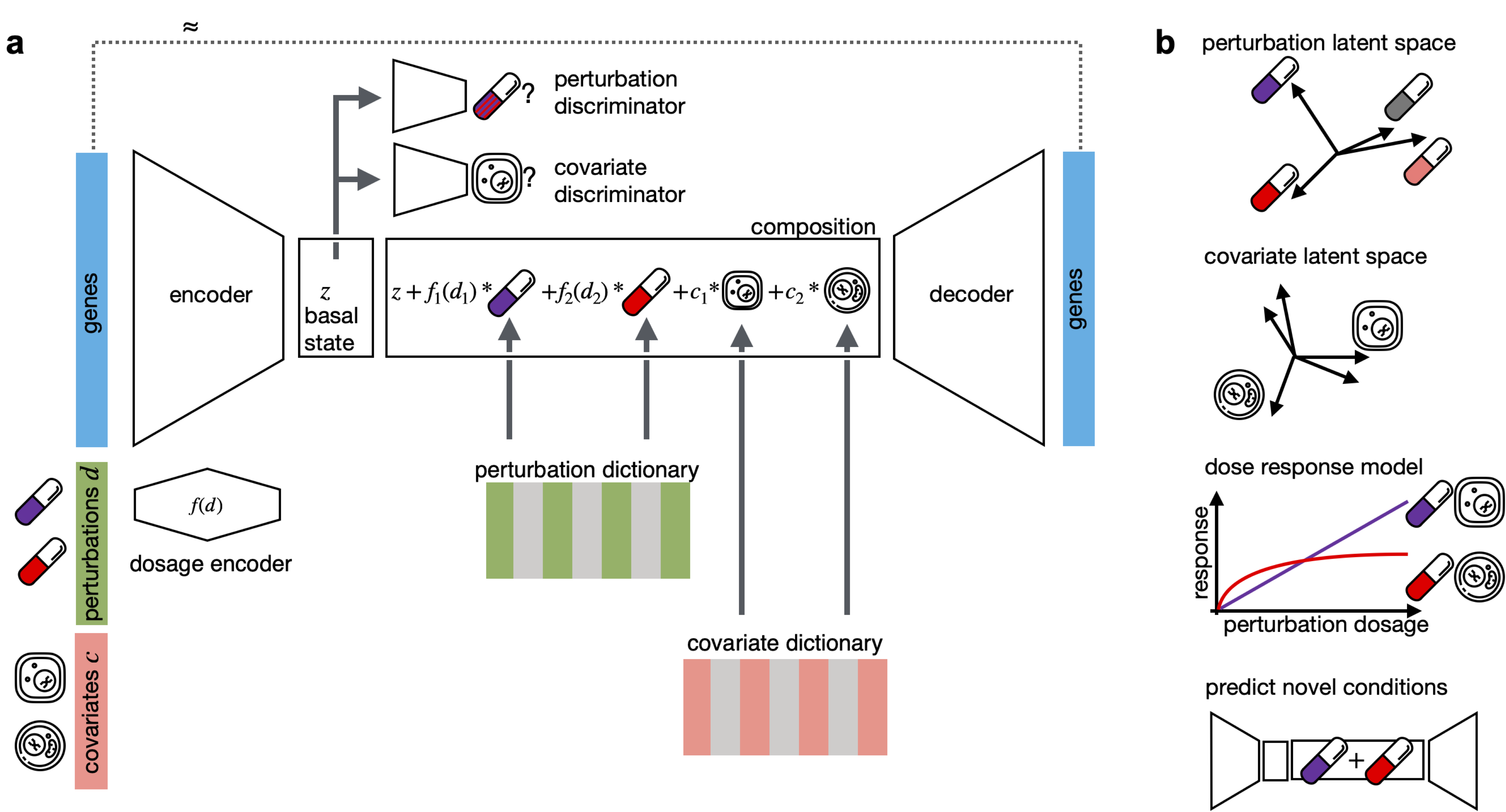
CPA is a framework to learn the effects of perturbations at the single-cell level. CPA encodes and learns phenotypic drug responses across different cell types, doses, and combinations. CPA allows:
- Out-of-distribution predictions of unseen drug combinations at various doses and among different cell types.
- Learn interpretable drug and cell-type latent spaces.
- Estimate the dose-response curve for each perturbation and their combinations.
- Access the uncertainty of the estimations of the model.
Usage and installation
See here for documentation and tutorials.
Support and contribute
If you have a question or new architecture or a model that could be integrated into our pipeline, you can post an issue
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
cpa-tools-0.7.0.tar.gz
(40.8 kB
view details)
Built Distribution
cpa_tools-0.7.0-py3-none-any.whl
(42.1 kB
view details)
File details
Details for the file cpa-tools-0.7.0.tar.gz.
File metadata
- Download URL: cpa-tools-0.7.0.tar.gz
- Upload date:
- Size: 40.8 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: poetry/1.1.13 CPython/3.8.13 Linux/5.4.0-132-generic
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 | 01400e1f11fbb437b3b0bed28bc4db14f64dc75189967bf407f21d5c7f1839ad |
|
| MD5 | 6d6b52356a6797b8766a2a9eb76a36d3 |
|
| BLAKE2b-256 | d81ad3a39bd1648c8e7a38c9baa8476f18723c665eb3d313f09af020424dae7a |
File details
Details for the file cpa_tools-0.7.0-py3-none-any.whl.
File metadata
- Download URL: cpa_tools-0.7.0-py3-none-any.whl
- Upload date:
- Size: 42.1 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: poetry/1.1.13 CPython/3.8.13 Linux/5.4.0-132-generic
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 | 6afff792a167dfe4b77e6b2345916191bcd0f85d005995d88f2ec1d4e4acd7f0 |
|
| MD5 | f708286cde0fbb22744e7f87eff121fd |
|
| BLAKE2b-256 | 5b8d768702c91185c5775d09998797959407405e27081aea441edab8fa699ab6 |