Active Learning Toolkit for Healthcare Imaging
Project description
MONAI Label
MONAI Label is a server-client system that facilitates interactive medical image annotation by using AI. It is an open-source and easy-to-install ecosystem that can run locally on a machine with single or multiple GPUs. Both server and client work on the same/different machine. It shares the same principles with MONAI.
 |
 |
 |
 |
Features
The codebase is currently under active development.
- Framework for developing and deploying MONAI Label Apps to train and infer AI models
- Compositional & portable APIs for ease of integration in existing workflows
- Customizable labelling app design for varying user expertise
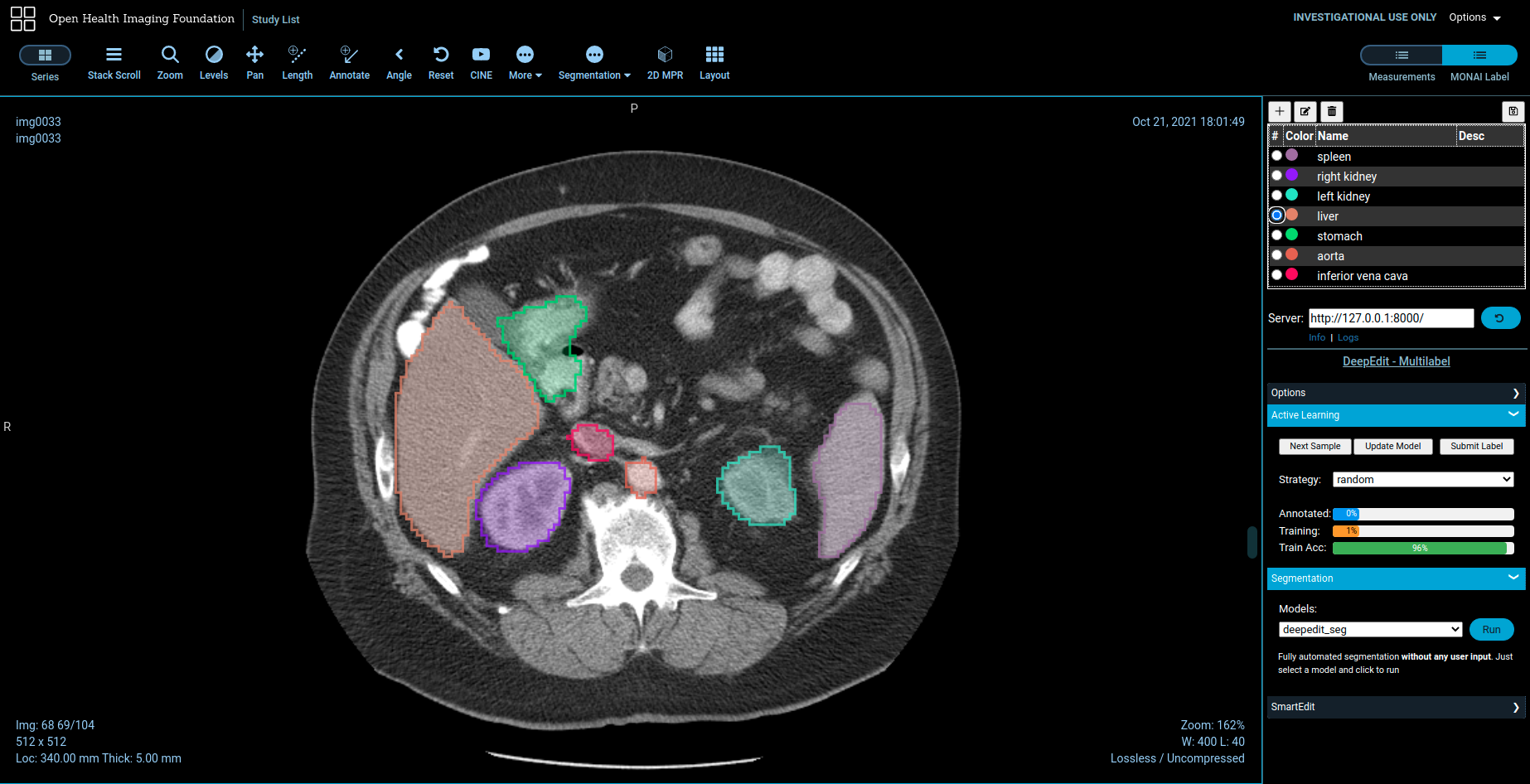
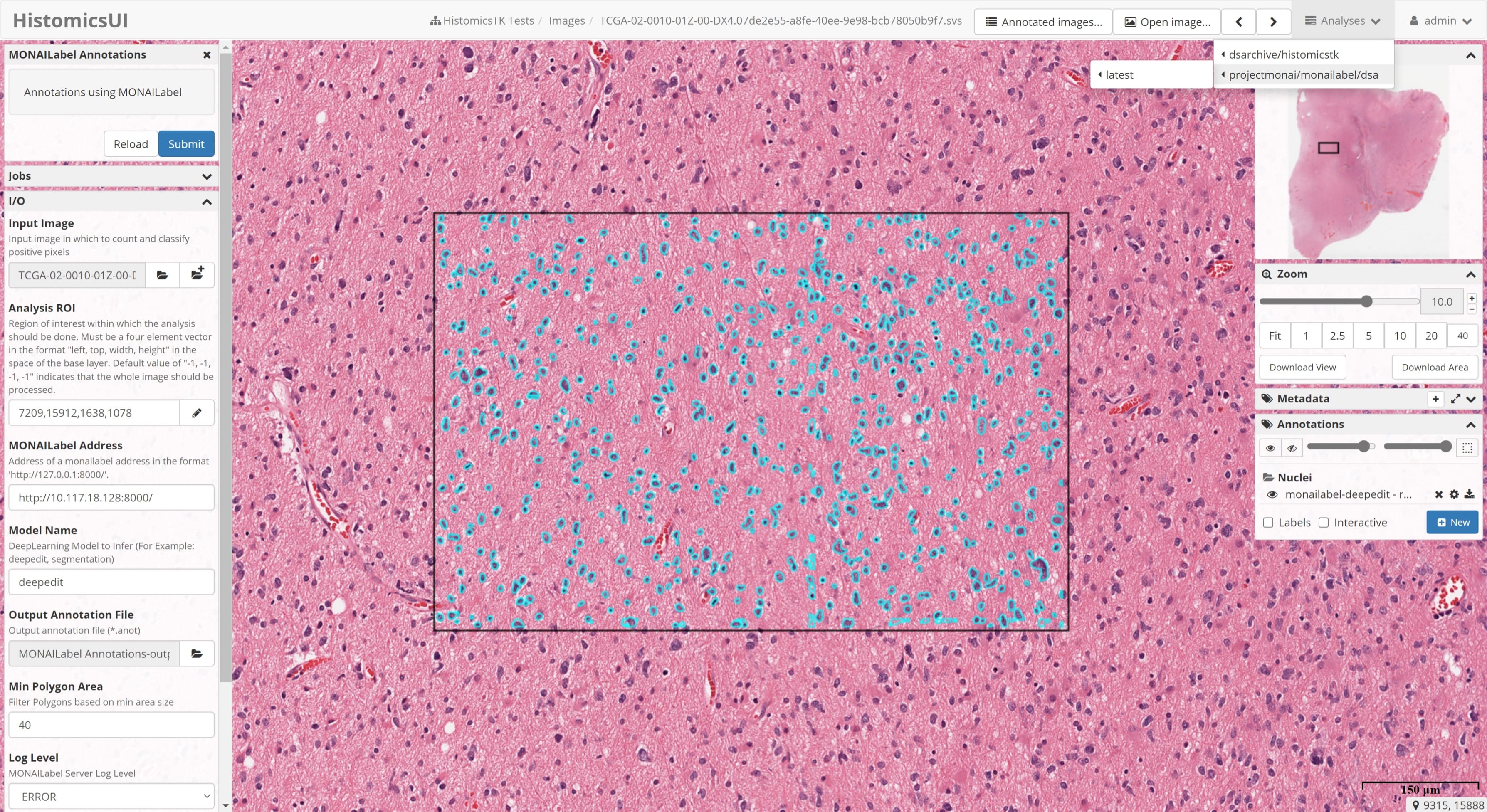
- Annotation support via 3DSlicer & OHIF for radiology
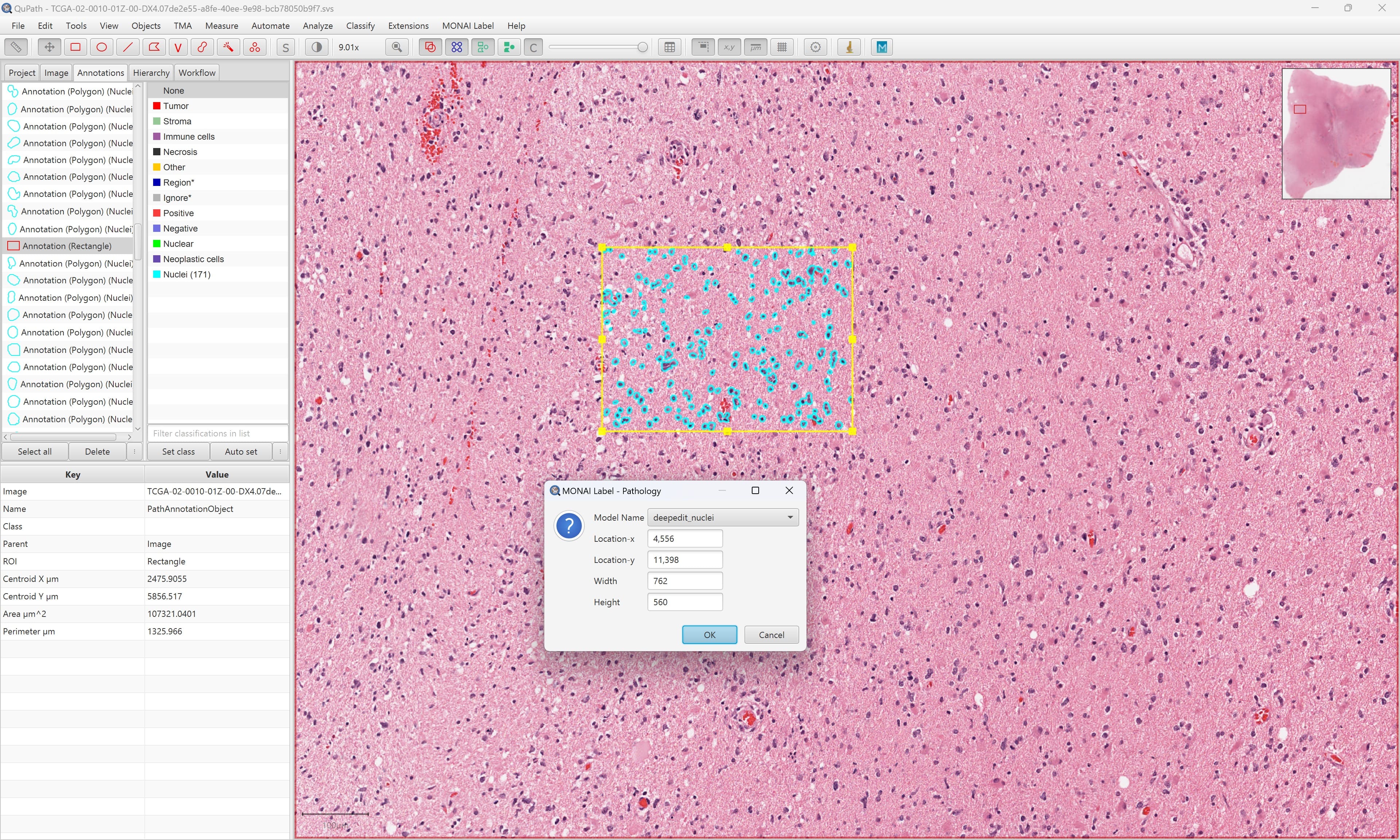
- Annotation support via QuPath , Digital Slide Archive & CVAT for pathology
- PACS connectivity via DICOMWeb
Installation
MONAI Label supports following OS with GPU/CUDA enabled.
- Ubuntu
- Windows
Current/Stable Version
pip install monailabel -U
Development version
To install the latest features using one of the following options:
Git Checkout (developer mode)
git clone https://github.com/Project-MONAI/MONAILabel
pip install -r MONAILabel/requirements.txt
export PATH=$PATH:`pwd`/MONAILabel/monailabel/scripts
Weekly Release
pip install monailabel-weekly -U
Docker
docker run --gpus all --rm -ti --ipc=host --net=host projectmonai/monailabel:latest bash
Once the package is installed, you can download sample radiology or pathology app and start monailabel server.
# download radiology app and sample dataset
monailabel apps --download --name radiology --output apps
monailabel datasets --download --name Task09_Spleen --output datasets
# start server using radiology app with deepedit model enabled
monailabel start_server --app apps/radiology --studies datasets/Task09_Spleen/imagesTr --conf models deepedit
More details refer docs: https://docs.monai.io/projects/label/en/stable/installation.html
If monailabel install path is not automatically determined, then you can provide explicit install path as:
monailabel apps --prefix ~/.local
For prerequisites, other installation methods (using the default GitHub branch, using Docker, etc.), please refer to the installation guide.
Once you start the MONAI Label Server, by default server will be up and serving at http://127.0.0.1:8000/. Open the serving URL in browser. It will provide you the list of Rest APIs available. For this, please make sure you use the HTTP protocol. You can provide ssl arguments to run server in HTTPS mode but this functionality is not fully verified across all clients.
Plugins
3D Slicer (radiology)
Download Preview Release from https://download.slicer.org/ and install MONAI Label plugin from Slicer Extension Manager.
Refer 3D Slicer plugin for other options to install and run MONAI Label plugin in 3D Slicer.
To avoid accidentally using an older Slicer version, you may want to uninstall any previously installed 3D Slicer package.
OHIF (radiology)
MONAI Label comes with pre-built plugin for OHIF Viewer. To use OHIF Viewer, you need to provide DICOMWeb instead of FileSystem as studies when you start the server.
Please install Orthanc before using OHIF Viewer. For Ubuntu 20.x, Orthanc can be installed as
apt-get install orthanc orthanc-dicomweb. However, you have to upgrade to latest version by following steps mentioned here.You can use PlastiMatch to convert NIFTI to DICOM
# start server using DICOMWeb
monailabel start_server --app apps/radiology --studies http://127.0.0.1:8042/dicom-web
OHIF Viewer will be accessible at http://127.0.0.1:8000/ohif/
NOTE: OHIF does not yet support Multi-Label interaction for DeepEdit. And you can still use 3D Slicer when MONAILabel is connected to DICOMWeb.
QuPath (pathology)
You can download sample whole slide images from https://portal.gdc.cancer.gov/repository
# start server using pathology over downloaded whole slide images
monailabel start_server --app apps/pathology --studies wsi_images
Refer QuPath for installing and running MONAILabel plugin in QuPath.
Digital Slide Archive (DSA) (pathology)
Refer Pathology for running a sample pathology use-case in MONAILabel.
NOTE: The DSA Plugin is under active development.
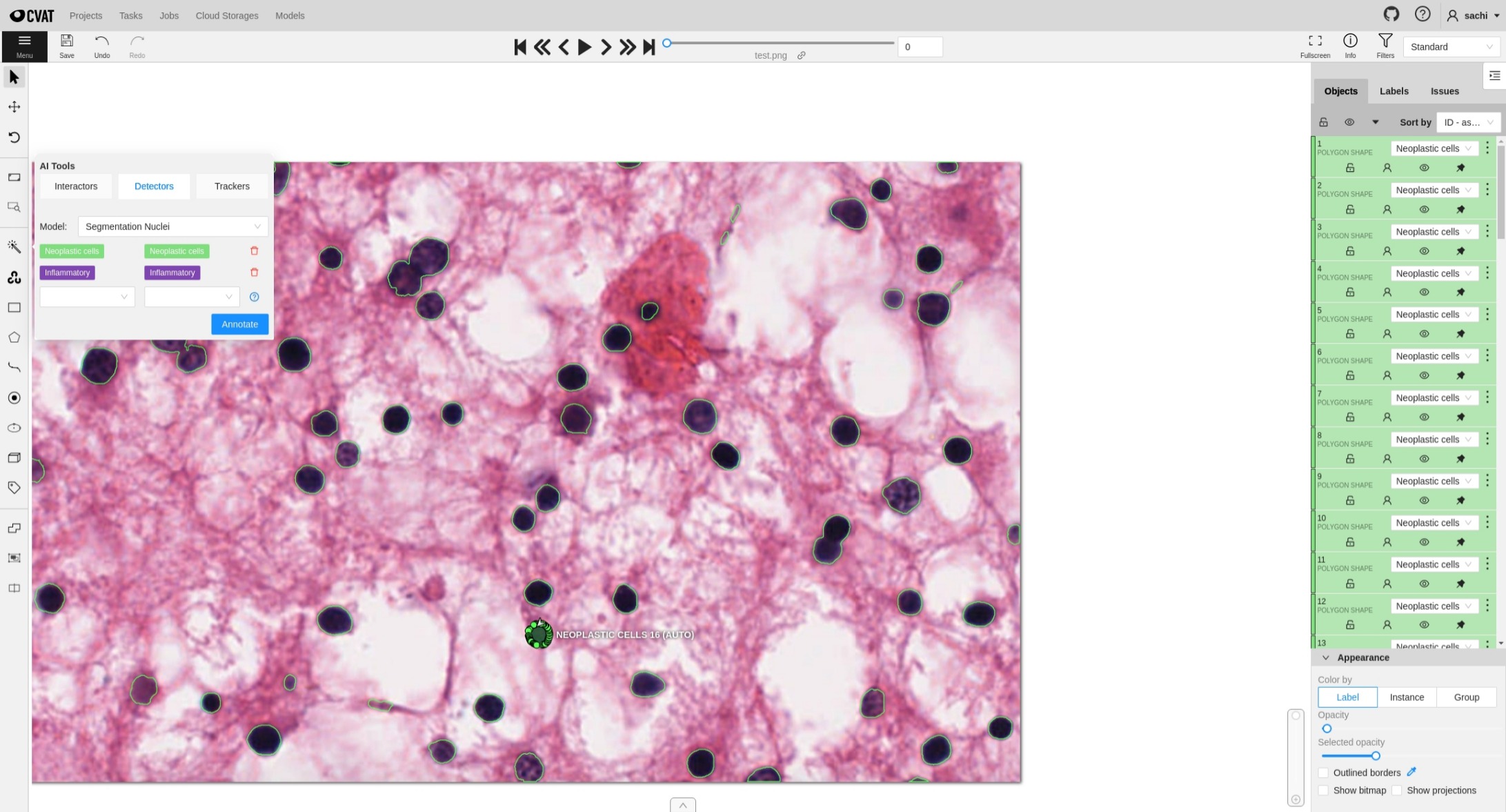
CVAT
Install CVAT and enable Semi-Automatic and Automatic Annotation . Refer CVAT Instructions for deploying available MONAILabel pathology models into CVAT.
Cite
If you are using MONAI Label in your research, please use the following citation:
@article{DiazPinto2022monailabel,
author = {Diaz-Pinto, Andres and Alle, Sachidanand and Ihsani, Alvin and Asad, Muhammad and
Nath, Vishwesh and P{\'e}rez-Garc{\'\i}a, Fernando and Mehta, Pritesh and
Li, Wenqi and Roth, Holger R. and Vercauteren, Tom and Xu, Daguang and
Dogra, Prerna and Ourselin, Sebastien and Feng, Andrew and Cardoso, M. Jorge},
title = {{MONAI Label: A framework for AI-assisted Interactive Labeling of 3D Medical Images}},
journal = {arXiv e-prints},
year = 2022,
url = {https://arxiv.org/pdf/2203.12362.pdf}
}
Contributing
For guidance on making a contribution to MONAI Label, see the contributing guidelines.
Community
Join the conversation on Twitter @ProjectMONAI or join our Slack channel.
Ask and answer questions over on MONAI Label's GitHub Discussions tab.
Links
- Website: https://monai.io/
- API documentation: https://docs.monai.io/projects/label
- Code: https://github.com/Project-MONAI/MONAILabel
- Project tracker: https://github.com/Project-MONAI/MONAILabel/projects
- Issue tracker: https://github.com/Project-MONAI/MONAILabel/issues
- Wiki: https://github.com/Project-MONAI/MONAILabel/wiki
- Test status: https://github.com/Project-MONAI/MONAILabel/actions
- PyPI package: https://pypi-hypernode.com/project/monailabel/
- Weekly previews: https://pypi-hypernode.com/project/monailabel-weekly/
- Docker Hub: https://hub.docker.com/r/projectmonai/monailabel
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distributions
Built Distribution
File details
Details for the file monailabel_weekly-0.4.dev2225-py3-none-any.whl.
File metadata
- Download URL: monailabel_weekly-0.4.dev2225-py3-none-any.whl
- Upload date:
- Size: 5.9 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.1 CPython/3.9.13
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 | a4196e7886e7f06205c0ed48a2436d47abeae904abbf8cc747a4b8629448f523 |
|
| MD5 | 7aec708891b20e3b6f310fe39b831b78 |
|
| BLAKE2b-256 | af5fff68a38a05c26daf154177d0cc21be8032850e554fee5b39023999081061 |