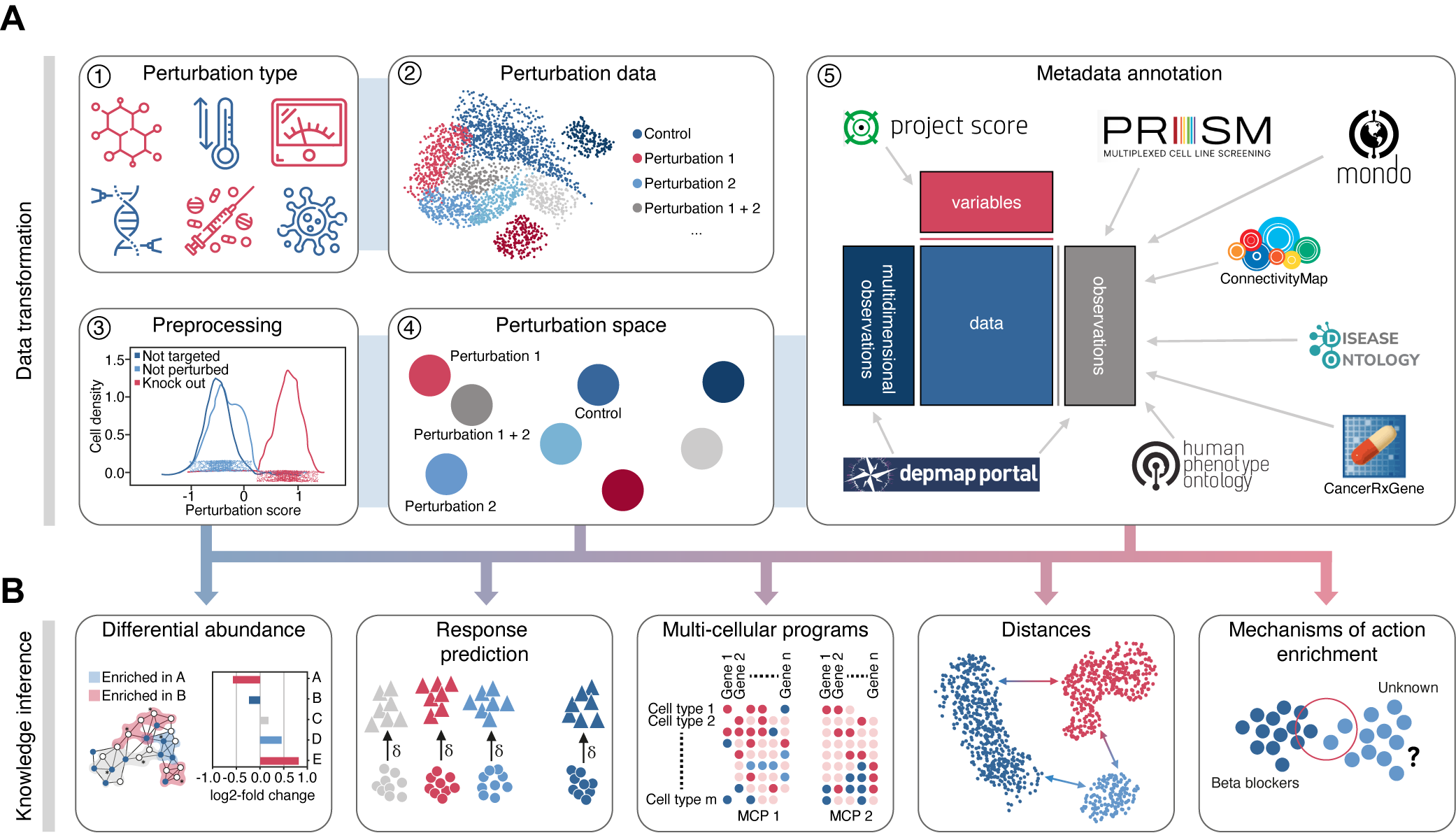
Perturbation Analysis in the scverse ecosystem.
Project description
pertpy
Documentation
Please read the documentation.
Installation
You can install pertpy via pip from PyPI:
pip install pertpy
if you want to use scCODA or tascCODA, please install pertpy as follows:
pip install pertpy[coda]
If you want to use the differential gene expression interface, please install pertpy by running:
pip install pertpy[de]
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
pertpy-0.8.0.tar.gz
(3.0 MB
view details)
Built Distribution
pertpy-0.8.0-py3-none-any.whl
(208.3 kB
view details)
File details
Details for the file pertpy-0.8.0.tar.gz.
File metadata
- Download URL: pertpy-0.8.0.tar.gz
- Upload date:
- Size: 3.0 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/5.1.0 CPython/3.12.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 | 2c0189cffc429f5737076fc47a9955fb6e59cb1ecbd8687a5bf66ddcc18abc89 |
|
| MD5 | 37d15bf06e17365947300ef44e16149b |
|
| BLAKE2b-256 | 621dca92308819eab559da710742416e7917ec663255bcb6ff5a478fc8b21ff9 |
Provenance
File details
Details for the file pertpy-0.8.0-py3-none-any.whl.
File metadata
- Download URL: pertpy-0.8.0-py3-none-any.whl
- Upload date:
- Size: 208.3 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/5.1.0 CPython/3.12.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 | fe410dc24172b9fd817f4734ee44ef4ceb7e91ac1702d90c33cbf0949c924008 |
|
| MD5 | 065180c01e1cc0efd02a2774c12cf202 |
|
| BLAKE2b-256 | d1cf48cc0a2d6c337d12cd14f60ac6bf378baa619764d37f5c991222ff13b2f9 |